United States (EN)
Select your region or country.


Polymerase chain reaction
(PCR)
Our photodetectors and imaging systems are used in PCR machines and equipment that require precise detection and analysis of fluorescence signals. PCR tests often involve the use of fluorescent dyes to detect the presence of the virus's genetic material. Our photonics technologies enhance the sensitivity and accuracy of these detection processes.
Fluorescence-based PCR tests rely on the detection of fluorescence signals emitted by labeled DNA fragments during the amplification process. Our photodetectors and imaging systems can be integrated into PCR instruments to capture and quantify these fluorescence signals, contributing to the accurate identification of the virus.
We provide instrumentation, such as high-quality cameras and detectors, that researchers and diagnostic labs can use to develop and improve PCR testing protocols. These tools can aid in optimizing the efficiency and reliability of PCR tests.
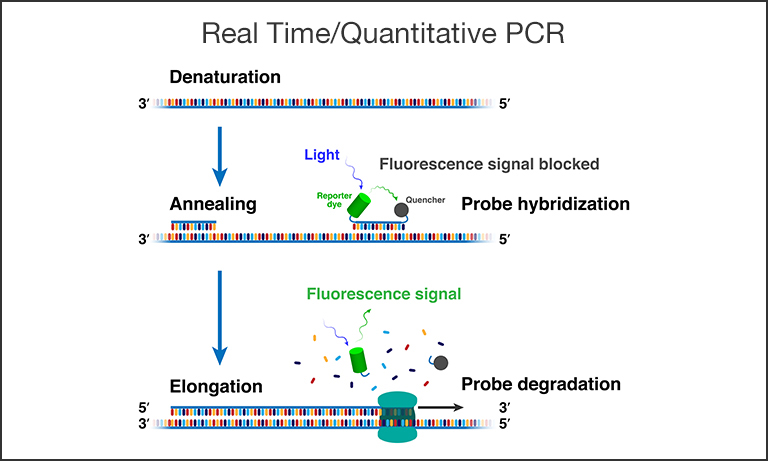
Figure 1. Schematic depiction of a single qPCR cycle. Shown above are: a) The denaturation stage when double-stranded DNA is separated into single strands (ssDNA) at high temperature. b) The annealing stage where the temperature is lowered, a primer binds to the three prime (3’) end of the specific sequence targeted for amplification, and (in the case of qPCR) a probe with a fluorescent reporter and quencher binds to the 5’ end. c) The elongation stage where the polymerase enzyme duplicates the sequence by attaching to the primer to start building a new complementary strand until it finishes by dissociating the probe and enabling the fluorescent signal.
Polymerase chain reaction, invented by Kary Mullis in 1983, has revolutionized not only the science of forensics but also the entire field of molecular biology and medicine by allowing us to chemically create billions of copies of a single DNA strand.
First, denaturation (Fig. 1) begins by heating a strand of DNA - mixed together with deoxyribonucleoside triphosphates (dNTPs), primers, Taq polymerase, and buffer - to a temperature of about 98℃ so that the two chains forming the double helix separate. The result is the formation of two DNA templates. In the next stage, called annealing, the temperature is lowered to 48℃-72℃, allowing DNA templates to combine with DNA primers (single-stranded nucleic acid).
DNA primers are short, single-stranded DNA sequences that attach at one end of each template and are very specific to the DNA intended for amplification. In the next stage, called extension or elongation, the temperature is about 68℃-72℃, and the primer recruits Taq polymerase to the site to add dNTPs to the templates forming replicas. These three steps are repeated, and the amplification is exponential so that the number of copies equals 2(N+1), where N is the number of cycles. If N=25, the number of copies is about 67 million.
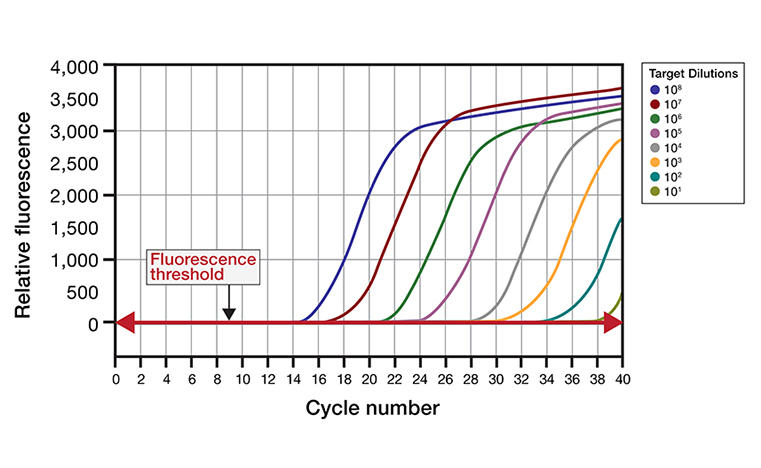
Figure 2. Amplification plot. Typical amplification charts of nucleic acid targets are generated by plotting the relative fluorescence signal detected by the sensor(s) as a function of the number of cycles. These charts usually include a series of dilutions in initial concentration of the amount of target added to the reaction. Once above the fluorescence threshold intensity, the signal increases exponentially then linearly as reaction reagents are consumed, finally reaching a plateau level.
A certain subset of PCR, real-time or quantitative PCR (qPCR), employs a fluorescent photonic technique called Forster resonance energy transfer (FRET) by adding an additional molecule into the mix, a probe (Fig. 1). Probes are short complementary sequences, also very specific to the intended target DNA, but with the addition of a fluorescent reporter dye and a FRET quencher in close proximity. Instead of binding at the beginning of the target DNA strand, the probe binds at the end, and, as the Taq polymerase finishes its replication, it cleaves the probe and destroys the FRET quenching relationship, allowing the fluorescent dye to report the duplicate strand to fluorescence detectors like cameras and silicon photodiodes. The resulting fluorescence intensity increases with the number of cycles in relation to the initial concentration of target in the sample (Fig. 2).
In this example of photonics applied to molecular biology, the quantity of DNA amplified can be monitored and detection of specific DNA or RNA needed for molecular diagnostic tests (Mol Dx) is possible. For example, testing for active infections in COVID-19 cases employs reverse transcriptase quantitative PCR (RT-qPCR). The virus is a positive-sense, single-stranded RNA (ribonucleic acid) type, using the RNA as the genetic material. Once inside a host, the virus uses its RNA to hijack the cell’s own protein machinery to disrupt normal cellular functions and replicate itself.
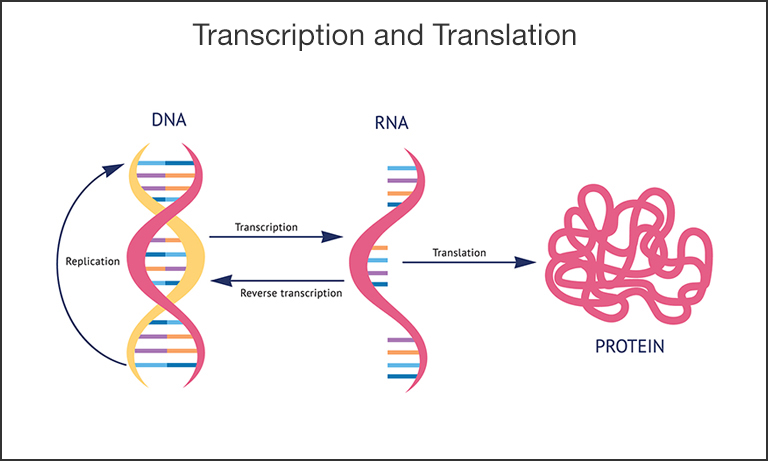
Figure 3. Schematic of the transcription and translation process. Encoded in DNA is the recipe for all biological function. The process of DNA replication occurs naturally, for example when cells divide, and can also be used as a tool to amplify the DNA. Transcription is the process in which the genetic code in DNA is read and transcribed into the complementary RNA molecule with the help of an enzyme, RNA polymerase. Different enzymes, called reverse transcriptases, can also invert this process and create DNA from RNA. Finally, translation is the process of creating proteins, the workhorse molecules in cells and organisms.
The reverse transcriptase term simply means the RNA is reverse transcribed (Fig. 3) back into complementary DNA (cDNA) prior to amplification by qPCR. Small quantities of viral genetic material in swab and saliva tests are undetectable even by the most sensitive photonic detectors, but when the genetic material is amplified millions of times, we can see the presence of the viral RNA. A key factor in the limit of detection (LOD) (Fig. 2) is more sensitive and wider dynamic range detectors, along with improvements in chemical reagents.
Lastly, digital PCR (dPCR) is yet another subset of this DNA amplification technology that employs photonic detectors to measure fluorescence similar to qPCR. However, in digital PCR, samples are split into thousands of reactions to detect and quantify exact copy numbers of the original targets. For example, dPCR is useful when true viral load counts or absolute quantities of specific genetic material need to be obtained. Due to the small volumes inherent in splitting a sample into thousands of separate reactions versus measuring the larger aggregate reaction, dPCR very often requires highly sensitive photodetectors, like silicon photomultipliers (SiPMs/MPPCs) and photomultiplier tubes (PMTs), capable of detecting even single photons from the fluorescent reporters.
Recommended products
Optics modules
The C16028-01 compact fluorescence detection optics module integrates excitation light sources and detectors in a sleek 60 × 60 × 22 mm design, allowing simultaneous measurement of FAM and ROX fluorescent dyes. With exceptional sensitivity down to 10^-10 W and independent LED control, it simplifies the construction of evaluation systems for precise and reliable fluorescence detection.
MPPC module C13365 series
MPPC modules are capable of detecting low-level light, made possible by their high internal gain, low dark noise and low crosstalk.

The small size MPPC with an effective area 1.3 mm x 1.3 mm has lower dark noise which enables more precise measurement.

The larger active area MPPC with an effective area of 3.0 mm x 3.00 mm has better light collection and a higher linear range of output.
Photosensor modules
These modules incorporate a high-sensitivity PMT for low light detection and an HVPS circuit.

This compact module integrates a photomultiplier tube, power supply, and amplifier, offering a highly sensitive detection solution with excellent signal amplification for low-light applications. Its plug-and-play design simplifies integration into systems for quick deployment. With a sensitivity wavelength range of 230-870 nm, this module features a multialkali photocathode.

Engineered for single-photon detection, this PMT delivers ultra-low noise performance, making it ideal for precise measurements in scientific research. The module integrates both the PMT power supply and biasing circuit for user-friendly operation.

This photosensor module features high-speed response and a wide dynamic range. It is optimized for time-critical applications like fluorescence lifetime measurements. With direct current output from the PMT, this module allows for photon counting and analog measurements. With a sensitivity wavelength range 230 to 920 nm, this module features an extended red multialkali photocathode.
Board level CMOS camera
This board captures 2D fluorescent images.

The C13752-50U digital CMOS board-level camera delivers high-resolution imaging with exceptional sensitivity in a compact, customizable form factor, making it ideal for OEM integration. Its fast frame rates and low power consumption provide efficient performance for industrial and scientific applications where space and power are limited.
- Confirmation
-
It looks like you're in the . If this is not your location, please select the correct region or country below.
You're headed to Hamamatsu Photonics website for US (English). If you want to view an other country's site, the optimized information will be provided by selecting options below.
In order to use this website comfortably, we use cookies. For cookie details please see our cookie policy.
- Cookie Policy
-
This website or its third-party tools use cookies, which are necessary to its functioning and required to achieve the purposes illustrated in this cookie policy. By closing the cookie warning banner, scrolling the page, clicking a link or continuing to browse otherwise, you agree to the use of cookies.
Hamamatsu uses cookies in order to enhance your experience on our website and ensure that our website functions.
You can visit this page at any time to learn more about cookies, get the most up to date information on how we use cookies and manage your cookie settings. We will not use cookies for any purpose other than the ones stated, but please note that we reserve the right to update our cookies.
1. What are cookies?
For modern websites to work according to visitor’s expectations, they need to collect certain basic information about visitors. To do this, a site will create small text files which are placed on visitor’s devices (computer or mobile) - these files are known as cookies when you access a website. Cookies are used in order to make websites function and work efficiently. Cookies are uniquely assigned to each visitor and can only be read by a web server in the domain that issued the cookie to the visitor. Cookies cannot be used to run programs or deliver viruses to a visitor’s device.
Cookies do various jobs which make the visitor’s experience of the internet much smoother and more interactive. For instance, cookies are used to remember the visitor’s preferences on sites they visit often, to remember language preference and to help navigate between pages more efficiently. Much, though not all, of the data collected is anonymous, though some of it is designed to detect browsing patterns and approximate geographical location to improve the visitor experience.
Certain type of cookies may require the data subject’s consent before storing them on the computer.
2. What are the different types of cookies?
This website uses two types of cookies:
- First party cookies. For our website, the first party cookies are controlled and maintained by Hamamatsu. No other parties have access to these cookies.
- Third party cookies. These cookies are implemented by organizations outside Hamamatsu. We do not have access to the data in these cookies, but we use these cookies to improve the overall website experience.
3. How do we use cookies?
This website uses cookies for following purposes:
- Certain cookies are necessary for our website to function. These are strictly necessary cookies and are required to enable website access, support navigation or provide relevant content. These cookies direct you to the correct region or country, and support security and ecommerce. Strictly necessary cookies also enforce your privacy preferences. Without these strictly necessary cookies, much of our website will not function.
- Analytics cookies are used to track website usage. This data enables us to improve our website usability, performance and website administration. In our analytics cookies, we do not store any personal identifying information.
- Functionality cookies. These are used to recognize you when you return to our website. This enables us to personalize our content for you, greet you by name and remember your preferences (for example, your choice of language or region).
- These cookies record your visit to our website, the pages you have visited and the links you have followed. We will use this information to make our website and the advertising displayed on it more relevant to your interests. We may also share this information with third parties for this purpose.
Cookies help us help you. Through the use of cookies, we learn what is important to our visitors and we develop and enhance website content and functionality to support your experience. Much of our website can be accessed if cookies are disabled, however certain website functions may not work. And, we believe your current and future visits will be enhanced if cookies are enabled.
4. Which cookies do we use?
There are two ways to manage cookie preferences.
- You can set your cookie preferences on your device or in your browser.
- You can set your cookie preferences at the website level.
If you don’t want to receive cookies, you can modify your browser so that it notifies you when cookies are sent to it or you can refuse cookies altogether. You can also delete cookies that have already been set.
If you wish to restrict or block web browser cookies which are set on your device then you can do this through your browser settings; the Help function within your browser should tell you how. Alternatively, you may wish to visit www.aboutcookies.org, which contains comprehensive information on how to do this on a wide variety of desktop browsers.
5. What are Internet tags and how do we use them with cookies?
Occasionally, we may use internet tags (also known as action tags, single-pixel GIFs, clear GIFs, invisible GIFs and 1-by-1 GIFs) at this site and may deploy these tags/cookies through a third-party advertising partner or a web analytical service partner which may be located and store the respective information (including your IP-address) in a foreign country. These tags/cookies are placed on both online advertisements that bring users to this site and on different pages of this site. We use this technology to measure the visitors' responses to our sites and the effectiveness of our advertising campaigns (including how many times a page is opened and which information is consulted) as well as to evaluate your use of this website. The third-party partner or the web analytical service partner may be able to collect data about visitors to our and other sites because of these internet tags/cookies, may compose reports regarding the website’s activity for us and may provide further services which are related to the use of the website and the internet. They may provide such information to other parties if there is a legal requirement that they do so, or if they hire the other parties to process information on their behalf.
If you would like more information about web tags and cookies associated with on-line advertising or to opt-out of third-party collection of this information, please visit the Network Advertising Initiative website http://www.networkadvertising.org.
6. Analytics and Advertisement Cookies
We use third-party cookies (such as Google Analytics) to track visitors on our website, to get reports about how visitors use the website and to inform, optimize and serve ads based on someone's past visits to our website.
You may opt-out of Google Analytics cookies by the websites provided by Google:
https://tools.google.com/dlpage/gaoptout?hl=en
As provided in this Privacy Policy (Article 5), you can learn more about opt-out cookies by the website provided by Network Advertising Initiative:
http://www.networkadvertising.org
We inform you that in such case you will not be able to wholly use all functions of our website.
Close


















































































































